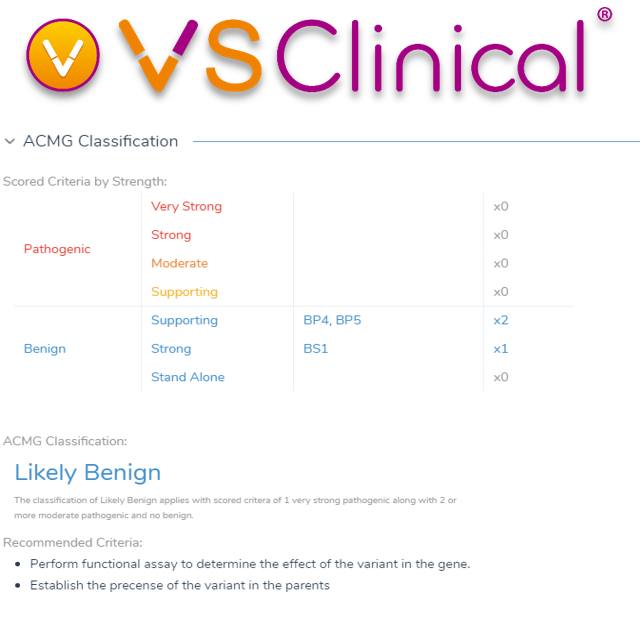
Variant Classification using ACMG/AMP Interpreting Sequence Guidelines - ClinGen | Clinical Genome Resource

ACMG classification of NRAS G12S. (Top) ProteinPaint display of somatic... | Download Scientific Diagram
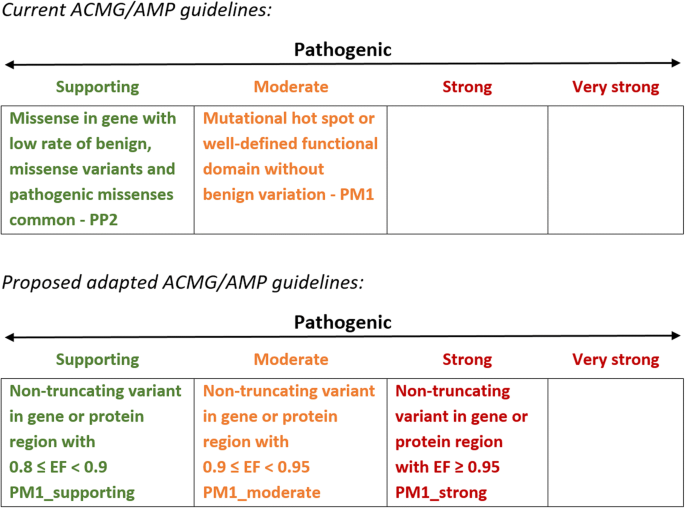
Quantitative approaches to variant classification increase the yield and precision of genetic testing in Mendelian diseases: the case of hypertrophic cardiomyopathy | Genome Medicine | Full Text

InterVar: Clinical Interpretation of Genetic Variants by the 2015 ACMG-AMP Guidelines. | Semantic Scholar
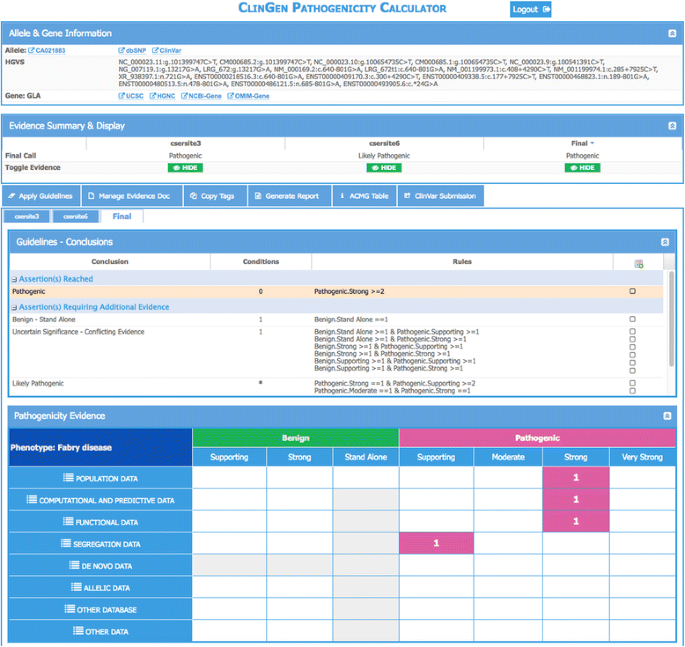
ClinGen Pathogenicity Calculator: a configurable system for assessing pathogenicity of genetic variants | Genome Medicine | Full Text

Cancer SIGVAR: A semi-automated interpretation tool for germline variants of hereditary cancer-related genes | bioRxiv

Current Tools, Databases, and Resources for Phenotype and Variant Analysis of Clinical Exome Sequencing - Advances in Molecular Pathology
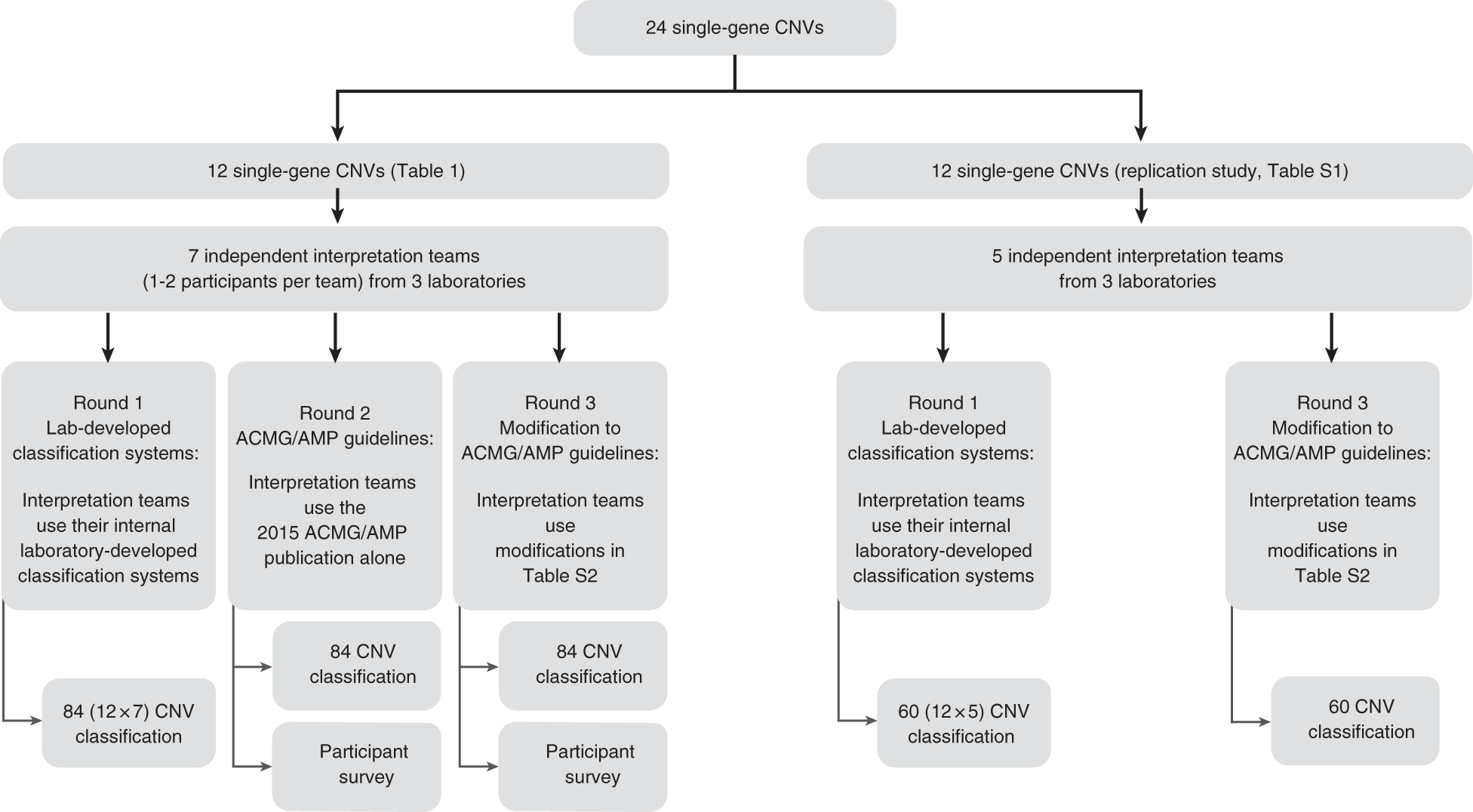
Adapting ACMG/AMP sequence variant classification guidelines for single-gene copy number variants | Genetics in Medicine

Classification and Reporting of Potentially Proarrhythmic Common Genetic Variation in Long QT Syndrome Genetic Testing | Circulation

Assessing performance of pathogenicity predictors using clinically relevant variant datasets | Journal of Medical Genetics

Quantifying the potential of functional evidence to reclassify variants of uncertain significance in the categorical and Bayesian interpretation frameworks. - Abstract - Europe PMC
Implementing ACMG guidelines on sequence variant interpretation: software-assisted variant curation and filtering
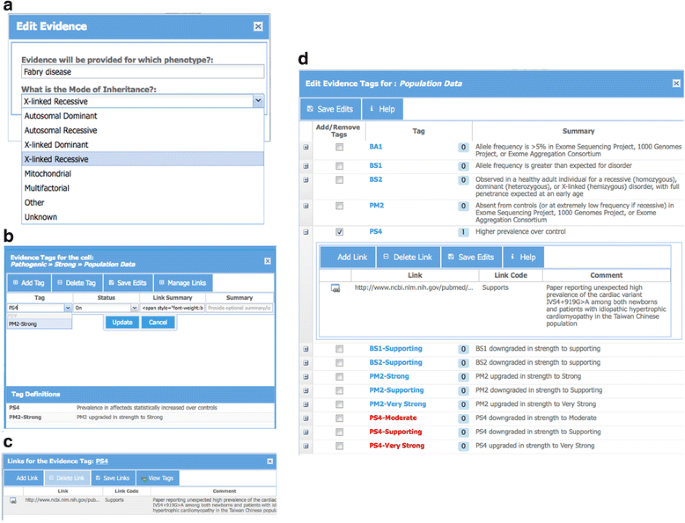
ClinGen Pathogenicity Calculator: a configurable system for assessing pathogenicity of genetic variants | Genome Medicine | Full Text
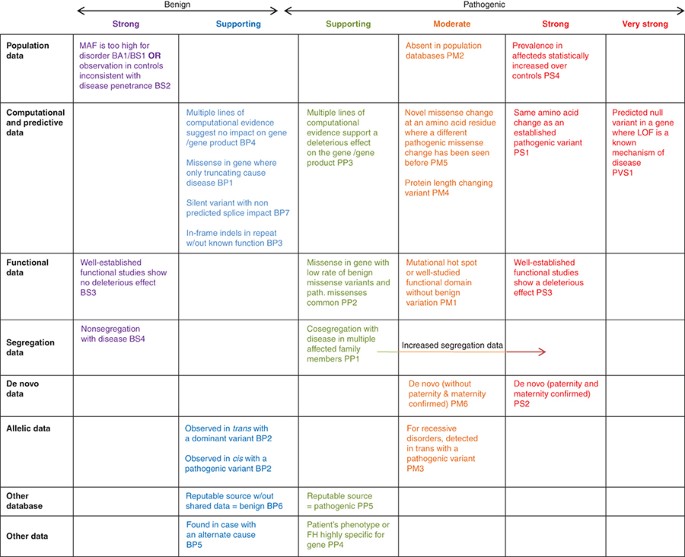



![PDF] ACGS Best Practice Guidelines for Variant Classification 2019 | Semantic Scholar PDF] ACGS Best Practice Guidelines for Variant Classification 2019 | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/86fa75f316823ef358a804fe1e45a98a85351bf9/4-Table1-1.png)